Among the various plastic recycling methods being explored, one promising approach is biological plastic recycling, also known as biorecycling, which utilizes enzymes or microorganisms to break down polymer molecules. One group of enzymes attracting attention is microbial cutinases. These enzymes are naturally produced by bacteria and fungi to degrade the waxy outer layer of plants, known as the cuticle. Because they can act on similar chemical bonds, they are considered promising for recycling poly(ethylene terephthalate) (PET), a plastic used in bottles and synthetic fibers.
However, applying these enzymes in industrial settings is not straightforward. PET is most efficiently degraded at temperatures around 70 °C, where it becomes more flexible and easier to process. At such high temperatures, enzymes must maintain a stable overall structure to avoid unfolding, while also retaining flexibility at their active site for molecular recognition and catalysis. This creates a design challenge, as structural rigidity and flexibility are often opposing properties.
To better understand this balance, a team of researchers led by Professor Tatsuya Nishino from the Department of Biological Science and Technology, Tokyo University of Science (TUS), Japan, along with Assistant Professor Sho Ito from the same department, and graduate researchers Mr. Ryohei Nojima (M.Sc., 2022) and Ms. Lirong Chen (M.Sc., 2024) from TUS, examined a heat-tolerant cutinase enzyme from the fungus Chaetomium thermophilum. The enzyme, known as CtCut, was analyzed under conditions relevant to high-temperature PET recycling to better understand how it maintains structural stability and catalytic potential. The study was published in Volume 16, Special Issue 4 of the journal Crystals on March 24, 2026.
“Plastic waste has become a severe problem in recent years, necessitating environmentally friendly recycling technologies. Thus, our aim was to contribute to the development of practical recycling technologies by clarifying the molecular basis of enzymes that function even under high-temperature conditions,” says Prof. Nishino.
For the study, the team created several versions of the enzyme. This included the wild-type (CtCutWT), which is the unmodified form, and a mutant version, CtCutS136A, in which the amino acid serine at position 136 is replaced with alanine.
They then determined the enzyme’s structure and assessed its thermal stability using differential scanning calorimetry, heating the protein from 30 °C to 100 °C to analyze how it absorbed heat.
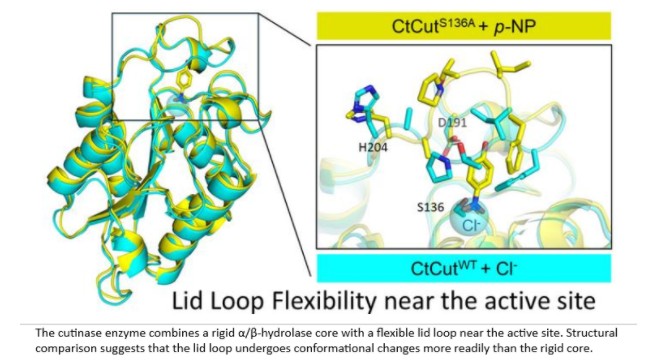
Structurally, the enzyme adopts a highly stable α/β-hydrolase fold, a common architecture among cutinases. Covering the active site is a flexible lid loop that can open and close. In its closed state, the active site is less accessible, but upon binding a molecule, the lid changes shape to allow binding and catalysis.
Notably, a chloride ion was found near the active site even when no substrate was present, suggesting that the active site forms a positively charged electrostatic microenvironment that may facilitate ligand binding.
As the enzyme was heated, it showed a two-step unfolding process, with a gradual transition beginning at around 60 °C, followed by a second transition near 65–70 °C. This indicates that different parts of the enzyme lose stability at different temperatures, suggesting the presence of structurally distinct regions within the protein.
“Our findings suggest the possibility of functional division within the enzyme. We observed that the mobile region near the active site undergoes structural changes in response to ligand binding, and that thermal denaturation proceeds in multiple stages,” says Prof. Nishino.
These findings support the idea that enzymes designed for plastic degradation may require both a stable overall structure and a flexible active site. The rigid core provides the thermal stability needed to withstand industrial conditions, while the flexible lid loop may help the enzyme adapt to bound molecules.
By better understanding this balance between stability and flexibility, the study provides new insights into the function of heat-tolerant enzymes and how they can be improved.
“Our study may lead to the development of technologies for efficiently decomposing and recycling PET in the future by providing design guidelines for enzymes that possess both heat resistance and potential catalytic capabilities for polymer degradation. This may address the growing challenge of plastic waste and help realize a sustainable resource-recycling society,” concludes Prof. Nishino.
Also Read:
Saturn’s magnetic bubble is lopsided compared to Earth’s
Scientists Find Hidden Critical Point in Water That Explains Strange Behavior