Mutated versions of the 80r and m396 antibodies can be produced and given as a therapeutic to fight the Covid-19 infection
A researcher, using computer models to understand the structure of viruses at the molecular level, has figured out how the old 2002 SARS coronavirus virus functions vis-a-vis the novel coronavirus or SARS-COV-2 that causes Covid-19. He discovered that sequence differences prevent 80R and m396 from binding to COVID-19 using ‘in silico’ analysis to fast-track passive immunity.
Since both illnesses (SARS-CoV and SARS-CoV-2) share the same spike protein, the entry key that allows the virus into the human cells, Padilla-Sanchez plans to take the antibodies found in the first outbreak in 2002 — 80R and m396 — and reengineer them to fit the current COVID-19 virus.
In his June 2020 publication in the online journal, Research Ideas and Outcomes, he describes efforts to unravel this problem using computer simulation based on his discovery that sequence differences prevent 80R and m396 from binding to COVID-19.
“Understanding why 80R and m396 did not bind to the SARS-CoV-2 spike protein could pave the way to engineering new antibodies that are effective,” Padilla-Sanchez said. “Mutated versions of the 80r and m396 antibodies can be produced and administered as a therapeutic to fight the disease and prevent infection.”
Supercomputing Process
His docking experiments showed that amino acid substitutions in 80R and m396 should increase binding interactions between the antibodies and SARS-CoV-2, providing new antibodies to neutralize the virus. “Now, I need to prove it in the lab,” he said.
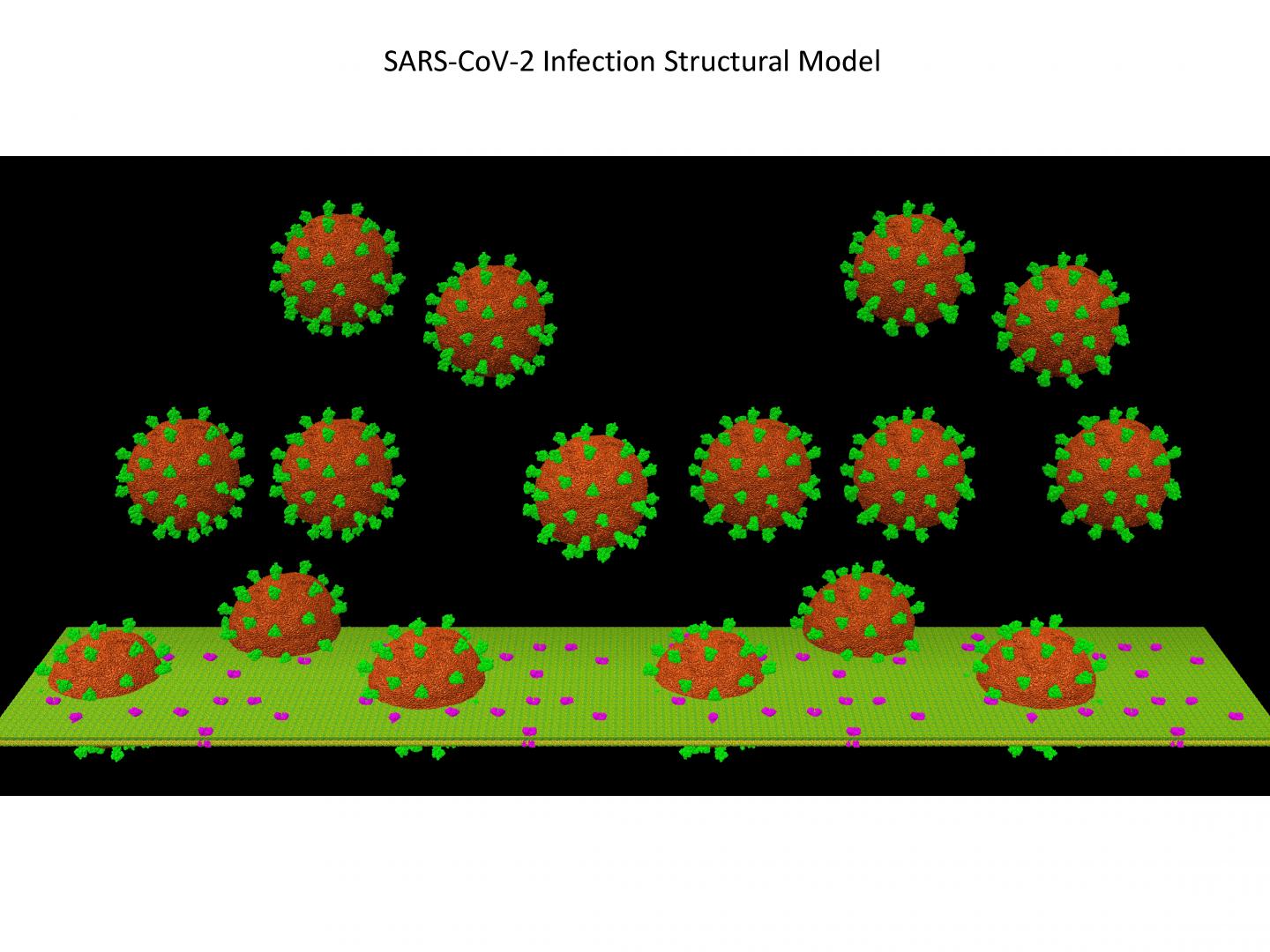
For this research, Padilla-Sanchez used supercomputing resources and ran the docking experiments for computational modeling and analysis of protein structures. The software virtually binds the proteins then provides a score for each binding experiment. “If you find a good docking position, then you can recommend that this new, mutated antibody should go to production.”
Currently, various labs across the world are already testing vaccines. “If we don’t find a vaccine in the near term we still have passive immunity, which can prevent infection for several months as long as you have the antibodies,” Padilla-Sanchez said. “Of course, a vaccine is the best outcome. However, passive immunity may be a fast track in providing relief for the pandemic.”
As of July 2020, COVID-19 has infected more than 16 million people worldwide with more than 630,000 deaths with no vaccine or therapeutics to fight the disease.

Pingback: Passive Immunity may be a fast track in providing relief for Covid-19 pandemic: Study | Medihelp Solutions